A subset of genes upregulated or downregulated by Cth2p under iron depletion. Genes exhibiting upregulation include HXK1 to YHR087W, while those undergoing downregulation comprise FIT1 to CCP1. FET3 serves as an example of a Cth2p-independent expression gene.
Abstract
Saccharomyces cerevisiae is widely used as a model organism for eukaryotic cells and generally prefers fermentation rather than respiration even under an aerobic environment. Only when glucose is exhausted, S. cerevisiae switches to aerobic respiration via massive reprogramming of gene expression accompanying that. These gene-expression changes are not simply achieved by the transcriptional level, rather multiple post-transcriptional regulatory steps are also involved. This chapter outlines how budding yeast cells coordinate energy metabolisms based on gene expression, with a focus on the intricate interplay of multiple post-transcriptional regulatory mechanisms. Especially, it includes the roles of RNA-binding proteins as well as non-coding RNAs for post-transcriptional regulations.
Keywords
- gene expression
- glucose
- fermentation
- respiration
- mitochondria
- post-transcription
- signaling
- RNA-binding protein
- non-coding RNA
1. Introduction
A fermentable sugar, glucose, is the preferred carbon and energy source, while concurrently serving as a signaling molecule [2, 3]. When yeasts are cultured in glucose-rich media under aerobic conditions, they predominantly employ glucose metabolism through glycolysis, yielding pyruvate as the primary product. Following glucose depletion, the fermented ethanol emerges as a carbon source, initiating a transition to respiration. This transformative metabolic process, known as a diauxic shift, unfolds in tandem with a substantial reconfiguration of gene expression, leading to dynamic upregulation including mitochondrial biogenesis and its functional capacities [4, 5, 6].
Glucose fermentation has superior catalytic efficiency compared to respiration in terms of adenosine triphosphate (ATP) production per unit of protein mass [7]. Conversely, respiration yields a tenfold increase in ATP per glucose molecule [8]. The equilibrium between respiration and fermentation is a pivotal determinant for unicellular survival [9]. Further, the oxidative fermentation or the Crabtree effect [10, 11], grants yeast an ecological advantage by allowing it to swiftly use glucose and produce ethanol, which possesses antiseptic properties [12].
This chapter overviews yeast metabolic systems, which are meticulously regulated through multiple steps at transcriptional and post-translational levels. It encompasses well-established signaling cascades and the regulation of nuclear-encoded mitochondrial gene expression, which interact with and are regulated by RNA-binding proteins and/or non-coding RNAs.
2. Metabolic arrangements: transitions between fermentation and respiration
Yeast has evolved efficient glucose utilization, employing both respiration and fermentation pathways to generate ATP from glucose. Both processes start with glycolysis, producing two molecules of pyruvate and ATP per glucose molecule. In the fermentation process, pyruvate is subsequently metabolized into ethanol. While this process does not yield additional ATP, it recycles nicotinamide adenine dinucleotide (NAD+) consumed during glycolysis, producing oxygen-independent ATP. In contrast, respiration involves the complete oxidation of pyruvate to CO2 through the tricarboxylic acid (TCA) cycle and oxidative phosphorylation (OXPHOS), yielding additional ATP in the presence of oxygen. Crabtree-positive yeasts, such as
3. Molecular and signaling aspects of yeast energy metabolic pathways
Glucose serves as a fundamental messenger molecule, playing a dual role as both an energy source and a signal for optimal growth conditions in cellular machinery. Yeasts, in particular, utilize glucose for this purpose. When the external glucose concentration exceeds 0.8 mM, yeast undergoes a transition into a mixed respiro-fermentative metabolism, resulting in ethanol production [17]. This shift underscores that the regulation between fermentation and respiration primarily corresponds to the sugar level [4]. Therefore, it is unsurprising that glucose plays a key element in shaping growth rate, fermentation capacity, and stress resistance [2, 18].
Multiple distinct pathways participate in responses to glucose. Some involve glucose interactions with cell surface receptors, while others require glucose import into the cell. These frequently utilized cascades can be classified into the following five signaling pathways: Ras-protein kinase A (PKA), Gpr1p–Gpa2p–PKA, the target of rapamycin (TOR)–Sch9p, Snf1p–Mig1p, or Snf3p–Rgt2p [3, 5, 19]. Among these, Ras and TOR, major global nutrient-sensing signal transduction cascades, serve pivotal roles in regulating cell growth in response to nutrient availability [18, 20]. Global glucose repression depends on an intracellular surge of cyclic adenosine monophosphate (cAMP), which activates PKA. Transcription rates significantly decrease upon Ras2p activation, which can occur independently of glucose presence and relies on a cAMP-responsive protein kinase [3, 21].
3.1 Ras–PKA and Gpr1p–Gpa2p–PKA
Ras, a guanine nucleotide-binding protein with seven transmembrane domains, activates adenylyl cyclase in its GTP-bound state. The addition of glucose to cells increases the level of GTP-bound Ras, resulting in an elevation of intracellular cAMP and subsequent activation of PKA [3, 21]. The PKA catalytic subunits, encoded by
Gpr1p, a plasma membrane protein (Figure 1), can sense the presence of glucose and/or sucrose and is coupled to Gα protein Gpa2p [23, 24]. This leads to the activation of adenylyl cyclase, resulting in an increase in cAMP concentration [2, 25, 26]. Subsequently, cAMP activates PKA. Therefore, it eventually controls the transcription of genes related to ribosome biogenesis and stress-responsive genes like
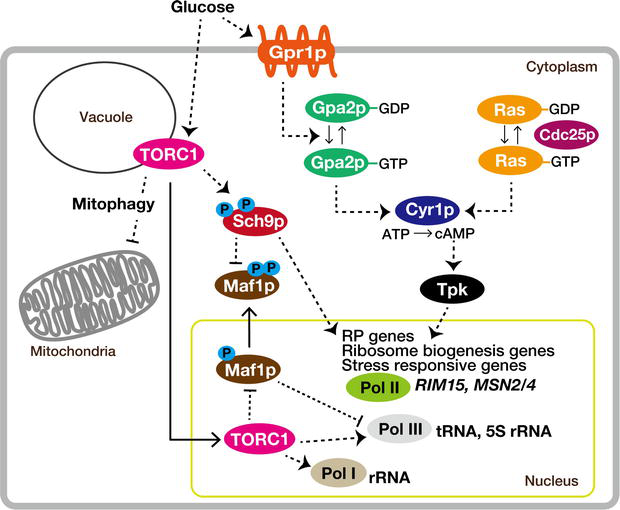
Figure 1.
Glucose signaling, facilitated by the small G-proteins Ras and Gpa2p, converges through PKA to stimulate ribosome biogenesis while concurrently suppressing the general stress response. Concurrently, within the TORC1 pathway, the kinase Sch9p plays a crucial role in strengthening the response of the PKA pathway. Dashed lines in the diagram symbolize regulatory interactions, which might not always be direct. See text for details.
3.2 TOR–Sch9p
TOR is a Ser/Thr kinase that was initially identified through yeast’s genetic screening [34]. The TOR proteins assemble into two structurally and functionally distinct complexes known as TOR complex 1 (TORC1) and TOR complex 2 (TORC2), of which only TORC1 is sensitive to rapamycin [35, 36]. The central components of the TOR consist of two TOR kinases paralogs, Tor1p, and Tor2p, along with a phosphate switch composed of the type 2A-related phosphatase Sit4p, TOR kinase phosphorylates Tap42p, and its inhibitor Tip41p [37, 38]. TORC1-dependent signals are mediated
The vacuolar surface primarily serves as the location for the TORC1 signaling pathway [40]. TORC1 is involved in respiration-induced mitophagy [41, 42]. This type of mitophagy is particularly important in cells that rely on OXPHOS for energy production, as disruptions in mitochondrial respiration can have significant consequences for cellular energy balance and overall cell function [43, 44].
An AGC family Ser/Thr kinase Sch9p is best characterized as the direct substrate of TORC1 [45]. Sch9p is a master regulator of ribosome biogenesis [45, 46, 47]. TORC1 controls all three RNA polymerase (Pol) systems
In contrast, gradual glucose exhaustion or abrupt withdrawal of glucose triggers a reduction in TORC1-dependent phosphorylation of five residues within the Sch9p C terminus [45, 57], leading to TORC1 inactivity. Inhibition of TORC1 provokes extensive transcriptome changes, reducing ribosomal particles by blocking the transcription of Pol I-dependent rRNA genes, Pol II-dependent RP genes, Pol III-dependent 5S rRNA, and the processing of 35S rRNA [18, 46, 47, 58]. Diminished phosphorylation of Sch9p transforms the Pol III repressor Maf1p from an inactivated to an active state [46, 59, 60]. This, in turn, leads to the suppression of Pol III transcriptome, including various small non-coding RNAs such as transfer RNA (tRNA). Consistent with TORC1’s functions, the absence of Tor1p results in an increase in mitochondrial respiration during glucose-based growth, primarily due to the enhanced translation of mitochondrial DNA (mtDNA)-encoded subunits of the OXPHOS complex. This effect is not observed in cells growing on glycerol [3, 61].
Recent research has shed light on the role of Snf1p–AMPK in fine-tuning TORC1 signaling during glucose starvation. Snf1p temporarily inhibits TORC1 activity by interacting with the phosphatidylinositol-3-phosphate (PI3P) and Kog1p-binding protein Pib2p [62]. This discovery highlights the mutual interaction between TOR and Snf1p, emphasizing their significance in metabolic adaptation.
3.3 Snf1p–Mig1p
The sucrose non-fermenting (Snf1) protein kinase, the yeast ortholog of mammalian AMP-activated S/T protein kinase (AMPK), is a central component of the primary glucose repression pathway responsible for adapting to glucose limitation [63, 64]. It forms a heterotrimeric complex with Snf4p (the regulatory γ-subunit) and one of the three β-subunits, Sip1p, Sip2p, or Gal83p, alongside the catalytic α-subunit, Snf1p [65]. Snf1p is activated under glucose limitation, assisting in energy homeostasis by promoting catabolic processes and inhibiting anabolic ones related to ATP generation and consumption [66]. This Snf1p complex regulates cellular processes through different transcription factors and enzymes [65]. For instance, Mig1p, a transcriptional repressor, controls the expression of genes involved in the metabolism and transportation of alternative carbon sources (e.g., maltose, galactose, sucrose) [67, 68]. Under glucose-rich conditions, Mig1p undergoes dephosphorylation and translocases to the nucleus (Figure 2). Together with the Ssn6p-Tup1p corepressor, it binds to target gene promoters [67, 69, 70]. Concurrently, Snf1p also undergoes dephosphorylation to prevent its nuclear localization in glucose-rich environments [71]. This dephosphorylation process involves protein phosphatase 1 (PP1), Glc7p–Reg1/2p, and possibly Sit4p, in collaboration with the phosphatase Ptc1p [72, 73, 74, 75, 76, 77].
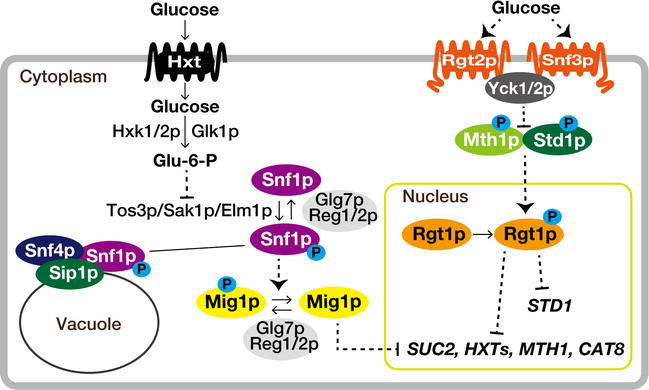
Figure 2.
The interconnected Snf and Rgt glucose signaling networks play a vital role in cellular regulation. The glucose sensor protein Snf3p, working alongside hexose transporters (Hxt), contributes to the yeast cell’s ability to sense extracellular glucose levels. Rgt1p, a transcription factor, governs glucose repression by controlling the expression of genes involved in glucose sensing and metabolism. Meanwhile, the Snf1p kinase and a transcriptional repressor Mig1p also respond to glucose availability. These components form a complex interplay in the yeast glucose signaling network, revealing the sophisticated regulatory mechanisms that govern cellular responses to glucose fluctuations.
3.4 Snf3p-Rgt2p
Two plasma membrane proteins, each composed of 12 transmembrane domains and featuring putative glucose-sensing capabilities, are Snf3p and Rgt2p (Figure 2). They share a resemblance to the Hxt glucose transporter [78], although they lack the capacity for glucose transport [79]. Their primarily role is regulating the expression of the seven main hexose transporters (HXT) genes [78].
Snf3p and Rgt2p act as glucose sensors, modulating the activity of Rgt1p, the transcription factor, in response to low and high glucose levels, respectively [80]. In the absence of glucose, Rgt1p, along with Ssn6p, Tup1p, Mth1p, and Std1p forms a repressor complex that inhibits
For more in-depth information on each cascade, comprehensive reviews are available [3, 18, 27, 79, 91].
4. Mitochondria: the central organelle of energy metabolism
Mitochondria, often referred to as the “powerhouses” of the cell, are essential organelles for both fermentation and respiration. At the diauxic shift, the mitochondrial volume expands concomitantly with the upregulation of Krebs cycle enzymes and respiratory complexes replete with abundant heme and Fe–S centers. Mitochondria are not discrete or autonomous entities; instead, they form highly dynamic and interconnected networks, and their biogenesis and structure are strongly influenced by the cell’s requirements [92, 93]. A classical targeting pathway for nuclear-encoded mitochondrial proteins uses mitochondrial targeting sequences (MTS) mainly located on their N-terminus [94, 95, 96], whereas approximately one-half of mRNAs for nuclear-encoded mitochondrial mRNAs are transported to the mitochondrial surface, and translated locally [97, 98, 99, 100]. Proximity-specific ribosome profiling targeting the tagged ribosomes on the mitochondrial surface showed highly enrichment of nuclear-encoded mitochondrial mRNAs, especially those encoding proteins in the mitochondrial inner membrane [101]. Cryo-electron cryotomography (CryoET) revealed active cytosolic ribosomes attach to the mitochondrial outer membrane and interact with the TOM complex [102]. The cytosolic translation of nuclear-encoded mitochondrial mRNAs and mitochondria indeed closely interact with each other.
4.1 Gene expression strategies for mitochondrial proteins
Mitochondria originated through the permanent integration of purple non-sulfur bacteria [103]. Throughout evolution, the majority of genes originally present in ancient bacteria have been transferred to nuclear DNA. Simultaneously, the genetic code within mitochondria has diverged from the conventional genetic code, resulting in significant differences in codon utilization between these two systems [104, 105]. In yeast today, mtDNA encodes only eight proteins, primarily associated with the OXPHOS system. Over 99% of mitochondrial proteins are instead encoded by the nuclear genome.
Under fermentable conditions, it is often assumed that nuclear-encoded mitochondrial genes are completely inactive due to subdued mitochondrial biogenesis and function. However, in reality, these nuclear-encoded mitochondrial genes remain transcriptionally active but subsequently undergo translational repression and/or rapid mRNA degradation [106, 107, 108]. Conversely, when exposed to respiratory conditions, there is a substantial upregulation of nuclear-encoded mitochondrial mRNAs, followed by increased translation to support mitochondrial biogenesis and enhance oxidative catabolism of carbon substrates. This orchestrated coordination between genomic and mitochondrial gene expression, along with the accurate sorting of nuclear-encoded mitochondrial proteins, is essential for maintaining optimal mitochondrial function [95, 109, 110, 111]. Disruptions in this process, such as the abnormal buildup of mitochondrial precursors in the cytosol leading to mitochondrial precursor over-accumulation stress (mPOS), or mitochondrial dysfunction, can activate a cytosolic proteostasis system [112, 113].
4.2 Iron: a vital element for mitochondrial function
Mitochondria continuously synthesize heme and Fe/S clusters while also facilitating amino acid and lipid metabolism [114, 115, 116]. Iron availability is essential for mitochondrial function and significantly impacts cellular metabolic responses to changes in carbon availability. Interestingly, yeast can survive OXPHOS defects and even complete loss of mtDNA but not disruption of mitochondrial Fe/S assembly, which proves to be fatal [114, 116, 117]. This is because the mitochondrial iron-sulfur cluster (ISC) assembly machinery is essential for the biogenesis of all cellular Fe/S proteins, including those in the cytosol and nucleus, which are involved in DNA maintenance and protein translation [114, 118].
Because iron is essential for cellular processes, when it is scarce, yeast employs Cth2p, an RNA-binding protein induced during iron starvation, to manage iron resources efficiently. Cth2p has a dual role: it suppresses non-essential iron consumption while promoting critical iron-dependent activities, including the assembly of ribonucleotide reductase (RNR) when iron is limited. Cth2p also inhibits mRNAs with AU-rich elements (ARE), mainly those related to iron metabolism and utilization, affecting pathways such as the TCA cycle, lipid biosynthesis, amino acid synthesis, and cofactor production. Additionally, Cth2p prevents excess iron accumulation in vacuoles by degrading mRNAs responsible for iron transport, including
| Gene | Function |
|---|---|
| Hexokinase isoenzyme 1 | |
| High-affinity glucose transporter | |
| High-affinity glucose transporter | |
| 6-phosphogluconolactonase | |
| Phosphoglucomutase; catalyzes the conversion from glucose-1-phosphate to glucose-6-phosphate | |
| Methylglyoxalase that converts methylglyoxal to D-lactate; involved in diauxic shift and stationary phase survival | |
| Haze-protective mannoprotein | |
| Glycogen phosphorylase required for the mobilization of glycogen | |
| Glycogen synthase; expression induced by glucose limitation | |
| Cytoplasmic protein that inhibits Gdb1p glycogen debranching activity | |
| Cytoplasmic aldehyde dehydrogenase | |
| Ornithine carbamoyltransferase | |
| Mitochondrial outer membrane protein | |
| Subunit Vb of cytochrome c oxidase | |
| Plasma membrane protein involved in maintaining membrane organization | |
| Endosomal protein involved in turnover of plasma membrane proteins | |
| Forms a complex with Rec102p and Spo11p necessary during the initiation of recombination | |
| Mutant is defective in directing meiotic recombination events to homologous chromatids | |
| Involved in RNA metabolism | |
| Cell wall mannoprotein involved in siderophore-Fe uptake | |
| Cell wall mannoprotein involved in siderophore-Fe uptake | |
| Heme binding peroxidase involved in reutilization of heme Fe | |
| Citrate synthase | |
| Mitochondrial isoform of citrate synthase | |
| Mitochondrial aconitase, Fe-S cluster protein | |
| Alpha-ketoglutarate dehydrogenase | |
| Dihydrolipoyl transsuccinylase | |
| Succinate dehydrogenase (ubiquinone) Fe-S cluster subunit | |
| Succinate dehydrogenase membrane anchor heme-binding subunit | |
| Subunit of cytochrome c oxidase | |
| Flavin-dependent monooxygenase, ubiquinone biosynthesis | |
| Ribonucleotide-diphosphate reductase | |
| Transporter that mediates vacuolar Fe storage | |
| RNase L inhibitor, Fe-S cluster protein | |
| NAD+-dependent glutamate synthase (GOGAT) | |
| Mitochondrial cytochrome-c peroxidase | |
| Ferro-O2-oxidoreductase |
Table 1.
Comprehensive reviews with more in-depth information mitochondrial protein sorting, iron homeostasis are available [114, 119, 120, 125, 126, 127].
5. Post-transcriptional gene expression regulation amid dynamic glucose changes
Gene expression is a multifaceted process that goes beyond mere transcriptional regulation, encompassing intricate post-transcriptional control mechanisms. It is not solely determined by transcriptional status; rather, it involves a complex interplay of factors. To optimize their growth conditions, cells undergo adaptations by adjusting their energy requirements through the modulation of vital metabolic enzymes, frequently accomplished
Glucose depletion triggers a swift and substantial halt in protein synthesis, which can be rapidly reversed upon glucose replenishment [137]. Additionally, this glucose depletion induces the formation of mRNA processing bodies (P-bodies), which act as central hubs where components of the 5–3′ mRNA decay pathway converge [138, 139, 140]. This compartmentalization of mRNAs in the cytosol potentially leads to translational repression and the degradation of specific mRNAs (although it has not been definitively proven). This phenomenon allows for a reduction in energy consumption while, at the same time, enabling the rapid translation of specific mRNAs. This facilitates the production of proteins necessary for adaptation [138, 141, 142, 143]. P-bodies indeed exclude translational machineries, including ribosomal components [144]. This specific response can significantly impact gene expression on a large scale. Further, during a diauxic shift, essential core components of P-bodies, Dhh1p and Pat1p, known for their roles as mRNA decapping activators and translational repressors, undergo a change in their intracellular localization. They shift from being excluded from polysomes in rapidly growing cells to co-localizing with polysomes [145].
Many aspects regarding P-bodies still remain obscure [143], but cells strategically employ adaptive mechanisms to dynamically regulate mRNA translation and degradation to manage the cellular protein repertoire. These processes are particularly important during glucose depletion and diauxic shifts. The formation of P-bodies and/or the dynamic behavior of core components such as Dhh1p and Pat1p would ensure the cell’s survival and growth in response to changing environmental conditions.
6. tRNA: a dynamic player of protein synthesis and cellular adaptation
tRNA is a classical small non-coding RNA present universally in living organisms. It plays a fundamental role in translation, along with the ribosome [146]. The primary function of tRNA, transferring amino acids into ribosomes, is to guarantee the precise integration of amino acids into proteins. Alternations in nutrient availability, such as shifts in glucose levels, can impact tRNA expression and their modifications, thereby exerting a profound influence on the efficiency of protein synthesis for a diverse array of proteins.
6.1 tRNA movement
In rapidly growing yeasts, tRNAs account for approximately 15% of the total cellular RNAs [147]. The availability of tRNAs positively correlates with codon utilization, influencing the usage of corresponding codons and
Yeast’s mtDNA encodes a complete set of tRNAs (24 tRNA species) required for mitochondrial translation within the organelle [105]. However, two cytosolic tRNAs, tRNALysCUU [154] and tRNAGln [155], are imported into mitochondria, potentially playing a role in stress response. In the case of tRNALysCUU, cells use a unique interaction mechanism with the mitochondrial outer membrane-attached glycolytic enzyme enolase [105]. This interaction induces a conformational change in tRNALysCUU, increasing its affinity for another protein factor, pre-mitochondrial lysyl-tRNA synthetase (preMsk1p), ultimately facilitating its co-import into the mitochondrial matrix [156]. Since the majority of mitochondrial proteins are encoded by the nuclear genome, cytosolic translation involving nuclear-encoded tRNAs significantly influences mitochondrial function. Mutants with impaired function of tRNAGlnUUG or tRNALysUUU exhibit inappropriate activation of various starvation responses during rapid growth. These responses involve the upregulation of genes related to glucose and nitrogen catabolism, along with premature and inadequate activation of autophagy. These effects can be alleviated by overexpressing tRNAGlnUUG or tRNALysUUU, which lack specific modifications [157].
6.2 Balancing act: tRNA dynamics in response to cellular stress
tRNAs are very dynamic, continually shuttling between the nucleus and cytosol throughout their entire life. Approximately one-fifth of total tRNA genes (61 of 275) in yeast, contain a single intron with variable lengths ranging from 14 to 60 nucleotides [149, 158]. Intriguingly, these introns consistently occupy the canonical position, precisely one nucleotide 3′ from the anticodon. These intron-containing tRNA genes transcribe precursor tRNA (pre-tRNA) with the intron sequence forming an A-I pair with anticodon nucleotides. This A-I interaction disrupts the crucial codon-anticodon binding during translation [146, 159, 160]. Thus, tRNA splicing is a vital process to address this issue, for intron-containing pre-tRNAs. However, unlike in mammals, yeast tRNA splicing occurs at the mitochondrial periphery due to the presence of the tRNA splicing SEN complex, which is located in the mitochondrial outer membrane [146, 161]. Consequently, intron-containing pre-tRNAs are transported to the mitochondrial membrane for splicing, and some are subsequently re-imported into the nucleus post-splicing [146].
In the case of mature tRNA, they also show bidirectional movement between the cytoplasm and nucleus [146]. This movement is tightly regulated and responsive to various cellular conditions and signals (Figure 3). In specific instances, mature tRNAs re-enter the nucleus following their cytoplasmic function. Although the reasons for this re-import are not always fully elucidated, it is possibly linked to quality control mechanisms or other regulatory processes [162]. This bidirectional trafficking of mature tRNAs allows cells to finely tune translation processes in response to changing conditions and to maintain the precision of protein synthesis. Thus, it emphasizes the dynamic nature of tRNA trafficking, which would contribute to the accurate and adaptable synthesis of proteins in the cell [146].
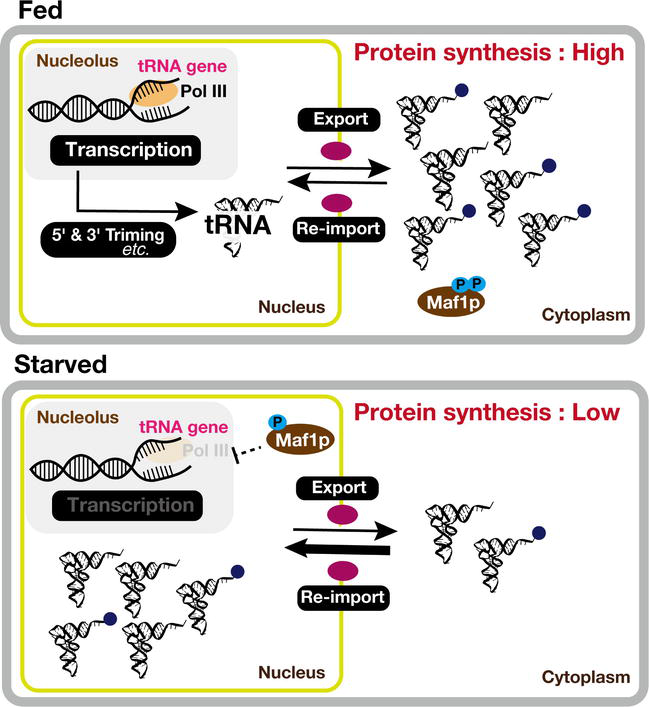
Figure 3.
Fine-tuning of tRNA dynamics in response to nutrient stress. tRNAs traverse between the nucleus and cytosol. This bidirectional movement is particularly crucial during varying cellular states, such as in nutrient-rich (fed) and nutrient-depleted (starved) conditions. Within the nucleus, tRNAs not only undergo transcription but also intricate maturation processes, including essential base modifications. Concurrently, in the cytosol, tRNAs actively engage in translation processes, fulfilling their indispensable role in protein synthesis. Under specific stress conditions, such as nutrient or glucose depletion, tRNAs are transported into the nucleus, deviating from cytosolic translation. This adaptive behavior, where tRNAs modulate both their localization and function in response to nutrient availability, underscores their critical involvement in cellular processes and protein synthesis
Remarkably, under glucose depletion, exposure to non-fermentable carbon sources such as glycerol, or in response to certain stressors, tRNA may accumulate in the nucleus [146, 163]. This sequestration of tRNA in the nucleus serves to isolate it from the cytoplasmic protein synthesis machinery, potentially serving as a mechanism to reduce protein synthesis globally [164, 165]. However, significant nuclear accumulation of cytoplasmic tRNAs does not necessarily result in a widespread inhibition of translation [166, 167]. Instead, the decrease in cytoplasmic tRNA levels during stress may involve the regulation of nuclear-cytoplasmic tRNA shuttling or changing tRNA transcription [165]. For instance, in
Comprehensive reviews with more in-depth information on tRNA are available [146, 169, 170, 171, 172].
7. Multifaceted regulator: rNA-binding protein Puf3p
Puf3p, a Pumilio homolog RNA-binding protein, is a well-known regulator of nuclear-encoded mitochondrial mRNAs [173, 174, 175]. Global analysis showed that Puf3p physically associates with 220 transcripts at least, and more than 70% of which are nuclear-encoded mitochondrial mRNAs [176]. Multiple multi-omics studies have consistently confirmed Puf3p’s binding specificity to nuclear-encoded mitochondrial mRNAs [177, 178, 179]. However, PAR-clip [178] and RIP-seq [179] have also identified numerous non-mitochondrial mRNAs as targets of Puf3p. Therefore, while Puf3p significantly influences the regulation of nuclear-encoded mitochondrial mRNAs, it also exerts a broader influence on gene expression associated with mitochondrial functions.
7.1 Molecular basis of Puf3p-RNA interaction: Structural analyzes and binding specificity determinants
Puf3p is comprised of eight Puf repeats, each composed of three α-helices, with neighboring repeats forming a crescent shape [180, 181, 182]. X-ray crystallography has revealed that three amino acid residues within each Puf repeat directly contact a single RNA base, determining binding specificity [182, 183, 184, 185, 186, 187, 188]. The Puf3p repeat domain (Puf3-RD) is sufficient to modulate mRNA metabolism and physically interacts with target mRNAs, exemplified by its binding to the 3′-UTR of
7.2 Multifaceted role of Puf3p in yeast physiology
Yeast Puf3p deletion mutants result in slow growth in respiratory media [176, 198], impair mitochondrial motility and biogenesis [198, 199], alter cellular oxidative stress tolerance and the glutathione redox state [200], and increase cellular oxygen consumption in a growth-dependent manner [201]. Under fermentation, Puf3p destabilizes its target mRNAs by promoting deadenylation and negatively regulates mitochondrial biogenesis [107, 108, 176, 190, 191, 201, 202, 203]. In agreement with its repressive roles in glucose-rich media, Puf3p’s abundance drastically decreases during the diauxic shift [199]. However, Puf3p associates with actively translating polysomes upon glucose depletion and promotes mitochondrial biogenesis [198, 204], indicating its bidirectional functions. These dual functions of Puf3p are regulated by phosphorylation
8. Conclusions
With its remarkable adaptability in metabolism and precision in gene regulation, yeast serves as a captivating model organism that holds significant implications for biotechnology and deepens our understanding of fundamental cellular intricacies. Its ability to expertly navigate the delicate equilibrium between glucose utilization, fermentation, and respiration underscores the core principles of cellular economics, which are vital for the survival and prosperity of all living organisms.
Furthermore, yeast’s intricate mechanisms for controlling post-transcriptional gene expression, involving processes such as mRNA processing, the dynamic behavior of tRNA, and the influence of RNA-binding proteins like Puf3p, exemplify an evolved strategy that enables cells to adapt to ever-changing environmental challenges rapidly. These dynamic processes serve as the linchpin for preserving cellular proteostasis, ensuring the precise and adaptable synthesis of proteins that play a pivotal role in sustaining and nurturing the growth of life.
Acknowledgments
This work was supported by JSPS KAKENHI Grant number JP20K06491 to S. H.
Abbreviations
adenosine triphosphate | |
nicotinamide adenine dinucleotide | |
tricarboxylic acid | |
oxidative phosphorylation | |
protein kinase A | |
target of rapamycin | |
cyclic adenosine monophosphate | |
TOR complex 1/2 | |
RNA polymerase | |
ribosomal protein | |
transfer RNA | |
mitochondrial DNA | |
phosphatidylinositol-3-phosphate | |
sucrose non-fermenting | |
AMP-activated S/T protein kinase | |
protein phosphatase 1 | |
hexose transporters | |
mitochondrial targeting sequences | |
cryo-electron cryotomography | |
mitochondrial precursor over-accumulation stress | |
iron-sulfur cluster | |
ribonucleotide reductase | |
AU-rich elements | |
mRNA processing bodies | |
precursor tRNA | |
Puf3p repeat domain |
References
- 1.
Tullio V. Yeast genomics and its applications in biotechnological processes: What is our present and near future? Journal of Fungi. 2022; 8 (7):752. DOI: 10.3390/jof8070752 - 2.
Rolland F, Winderickx J, Thevelein JM. Glucose-sensing and -signalling mechanisms in yeast. FEMS Yeast Research. 2002; 2 (2):183-201. DOI: 10.1073/pnas.0305901101 - 3.
Gancedo JM. The early steps of glucose signalling in yeast. FEMS Microbiology Reviews. 2008; 32 (4):673-704. DOI: 10.1111/j.1574-6976.2008.00117.x - 4.
Otterstedt K, Larsson C, Bill RM, Ståhlberg A, Boles E, Hohmann S, et al. Switching the mode of metabolism in the yeast Saccharomyces cerevisiae . EMBO Reports. 2004;5 (5):532-537. DOI: 10.1038/sj.embor.7400132 - 5.
Zaman S, Lippman SI, Zhao X, Broach JR. How Saccharomyces responds to nutrients. Annual Review of Genetics. 2008;42 :27-81. DOI: 10.1146/annurev.genet.41.110306.130206 - 6.
De Virgilio C. The essence of yeast quiescence. FEMS Microbiology Reviews. 2012; 36 (2):306-339. DOI: 10.1111/j.1574-6976.2011.00287.x - 7.
Nilsson A, Nielsen J. Metabolic trade-offs in yeast are caused by F1F0-ATP synthase. Scientific Reports. 2016; 6 :22264. DOI: 10.1038/srep22264 - 8.
van Dijken J, Bauer J, Brambilla L, Duboc P, Francois J, Gancedo C, et al. An interlaboratory comparison of physiological and genetic properties of four Saccharomyces cerevisiae strains. Enzyme and Microbial Technology. 2000;26 (9-10):706-714. DOI: 10.1016/s0141-0229(00)00162-9 - 9.
Molenaar D, Van Berlo R, De Ridder D, Teusink B. Shifts in growth strategies reflect tradeoffs in cellular economics. Molecular Systems Biology. 2009; 5 :323. DOI: 10.1038/msb.2009.82 - 10.
Crabtree HG. Observations on the carbohydrate metabolism of tumours. The Biochemical Journal. 1929; 23 (3):536-545. DOI: 10.1042/bj0230536%0A - 11.
Pfeiffer T, Morley A. An evolutionary perspective on the Crabtree effect. Frontiers in Molecular Biosciences. 2014; 1 :17. DOI: 10.3389/fmolb.2014.00017 - 12.
Dai Z, Huang M, Chen Y, Siewers V, Nielsen J. Global rewiring of cellular metabolism renders Saccharomyces cerevisiae Crabtree negative. Nature Communications. 2018;9 (1):3059. DOI: 10.1038/s41467-018-05409-9 - 13.
Piskur J, Rozpedowska E, Polakova S, Merico A, Compagno C. How did Saccharomyces evolve to become a good brewer? Trends in Genetics. 2006;22 (4):183-186. DOI: 10.1016/j.tig.2006.02.002 - 14.
Scott M, Gunderson CW, Mateescu EM, Zhang Z, Hwa T. Interdependence of cell growth. Science. 2010; 330 (6007):1099-1102. DOI: 10.1126/science.1192588 - 15.
Kussell E. Evolution in microbes. Annual Review of Biophysics. 2013; 42 (1):493-514. DOI: 10.1146/annurev-biophys-083012-130320 - 16.
Warburg O. On the origin of cancer. Science. 1956; 123 :309-314. DOI: 10.1126/science.123.3191.309 - 17.
Verduyn C, Zomerdijk TPL, van Dijken JP, Scheffers WA. Continuous measurement of ethanol production by aerobic yeast suspensions with an enzyme electrode. Applied Microbiology and Biotechnology. 1984; 19 (3):181-185. DOI: 10.1007/BF00256451 - 18.
Broach JR. Nutritional control of growth and development in yeast. Genetics. 2012; 192 (1):73-105. DOI: 10.1534/genetics.111.135731 - 19.
Santangelo GM. Glucose signaling in Saccharomyces cerevisiae . Microbiology and Molecular Biology Reviews. 2006;70 (1):253-282. DOI: 10.1128/MMBR.70.1.253-282.2006 - 20.
Schmelzle T, Beck T, Martin DE, Hall MN. Activation of the RAS/cyclic AMP pathway suppresses a TOR deficiency in yeast. Molecular and Cellular Biology. 2004; 24 (1):338-351. DOI: 10.1128/MCB.24.1.338-351.2004 - 21.
Tamanoi F. Ras signaling in yeast. Genes & Cancer. 2011; 2 (3):210-215. DOI: 10.1177/1947601911407322 - 22.
Wang Y, Pierce M, Schneper L, Güldal CG, Zhang X, Tavazoie S, et al. Ras and Gpa2 mediate one branch of a redundant glucose signaling pathway in yeast. PLoS Biology. 2004; 2 (5):610-622. DOI: 10.1371/journal.pbio.0020128 - 23.
Yun C-W, Tamaki H, Nakayama R, Yamamoto K, Kumagai H. G-protein coupled receptor from yeast Saccharomyces cerevisiae . Biochemical and Biophysical Research Communications. 1997;240 (2):287-292. DOI: 10.1006/bbrc.1997.7649 - 24.
Xue Y, Batlle M, Hirsch JP. GPR1 encodes a putative G protein-coupled receptor that associates with the Gpa2p Gα subunit and functions in a Ras-independent pathway. The EMBO Journal. 1998; 17 (7):1996-2007. DOI: 10.1093/emboj/17.7.1996 - 25.
Yun C-W, Tamaki H, Nakayama R, Yamamoto K, Kumagai H. Gpr1p, a putative G-protein coupled receptor, regulates glucose-dependent cellular cAMP level in yeast Saccharomyces cerevisiae . Biochemical and Biophysical Research Communications. 1998;252 (1):29-33. DOI: 10.1006/bbrc.1998.9600 - 26.
Kraakman L, Lemaire K, Ma P, Teunlssen AW, Donaton MC, Van Dijck P, et al. A Saccharomyces cerevisiae G-protein coupled receptor, Gpr1, is specifically required for glucose activation of the cAMP pathway during the transition to growth on glucose. Molecular Microbiology. 1999;32 (5):1002-1012. DOI: 10.1046/j.1365-2958.1999.01413.x - 27.
Conrad M, Schothorst J, Kankipati HN, Van Zeebroeck G, Rubio-Texeira M, Thevelein JM. Nutrient sensing and signaling in the yeast Saccharomyces cerevisiae . FEMS Microbiology Reviews. 2014;38 (2):254-299. DOI: 10.15698/mic2021.01.740 - 28.
Wera S, De Schrijver E, Geyskens I, Nwaka S, Thevelein JM. Opposite roles of trehalase activity in heat-shock recovery and heat-shock survival in Saccharomyces cerevisiae . The Biochemical Journal. 1999;343 (3):621-626 - 29.
François JM, Walther T, Parrou JL. Genetics and regulation of glycogen and trehalose metabolism in Saccharomyces cerevisiae . In: Systems Biology. New York: SpringerLink; 2012. pp. 29-55. DOI: 10.1007/978-3-642-21467-7 - 30.
Wingender-Drissen R, Becker JU. Regulation of yeast phosphorylase by phosphorylase kinase and cAMP-dependent protein kinase. FEBS Letters. 1983; 163 (1):33-36. DOI: 10.1016/0014-5793(83)81156-9 - 31.
Dihazi H, Kessler R, Eschrich K. Glucose-induced stimulation of the Ras-cAMP pathway in yeast leads to multiple phosphorylations and activation of 6-phosphofructo-2-kinase. Biochemistry. 2003; 42 (20):6275-6282. DOI: 10.1021/bi034167r - 32.
Portela P, Moreno S, Rossi S. Characterization of yeast pyruvate kinase 1 as a protein kinase A substrate, and specificity of the phosphorylation site sequence in the whole protein. The Biochemical Journal. 2006; 396 (1):117-126. DOI: 10.1042/BJ20051642 - 33.
Mazón MJ, Gancedo JM, Gancedo C. Inactivation of yeast fructose-1,6-bisphosphatase. In vivo phosphorylation of the enzyme. The Journal of Biological Chemistry. 1982; 257 (3):1128-1130 - 34.
Heitman J, Movva NR, Hall MN. Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast. Science. 1991; 253 (5022):905-909. DOI: 10.1126/science.1715094 DOI: 10.1126/science.1715094 - 35.
González A, Hall MN. Nutrient sensing and TOR signaling in yeast and mammals. The EMBO Journal. 2017; 36 (4):397-408. DOI: 10.15252/embj.201696010 - 36.
Loewith R, Jacinto E, Wullschleger S, Lorberg A, Crespo JL, Bonenfant D, et al. Two TOR complexes, only one of which is rapamycin sensitive, have distinct roles in cell growth control. Molecular Cell. 2002; 10 (3):457-468. DOI: 10.1016/s1097-2765(02)00636-6 - 37.
Düvel K, Broach JR. The role of phosphatases in TOR signaling in yeast. In: Current Topics in Microbiology and Immunology. New York: SpringerLink; 2004. pp. 19-38. DOI: 10.1007/978-3-642-18930-2_2 - 38.
Jacinto E, Guo B, Arndt KT, Schmelzle T, Hall MN. TIP41 interacts with TAP42 and negatively regulates the TOR signaling pathway. Molecular Cell. 2001; 8 (5):1017-1026. DOI: 10.1016/s1097-2765(01)00386-0 - 39.
Breitkreutz A, Choi H, Sharom JR, Boucher L, Neduva V, Larsen B, et al. A global protein kinase and phosphatase interaction network in yeast. Science. 2010; 328 (5981):1043-1046. DOI: 10.1126/science.1176495 - 40.
Sturgill TW, Cohen A, Diefenbacher M, Trautwein M, Martin DE, Hall MN. TOR1 and TOR2 have distinct locations in live cells. Eukaryotic Cell. 2008; 7 (10):1819-1830. DOI: 10.1128/EC.00088-08 - 41.
Liu Y, Okamoto K. The TORC1 signaling pathway regulates respiration-induced mitophagy in yeast. Biochemical and Biophysical Research Communications. 2018; 502 (1):76-83. DOI: 10.1016/j.bbrc.2018.05.123 - 42.
Innokentev A, Kanki T. Mitophagy in yeast: Molecular mechanism and regulation. Cell. 2021; 10 (12):3569. DOI: 10.3390/cells10123569 - 43.
May AI, Prescott M, Ohsumi Y. Autophagy facilitates adaptation of budding yeast to respiratory growth by recycling serine for one-carbon metabolism. Nature Communications. 2020; 11 (1):5052. DOI: 10.1038/s41467-020-18805-x - 44.
Kumar R, Reichert AS. Autophagy promotes mitochondrial respiration by providing serine for one-carbon-metabolism. Autophagy. 2021; 17 (12):4480-4483. DOI: 10.1080/15548627.2021.1909408 - 45.
Urban J, Soulard A, Huber A, Lippman S, Mukhopadhyay D, Deloche O, et al. Sch9 is a major target of TORC1 in Saccharomyces cerevisiae . Molecular Cell. 2007;26 (5):663-674. DOI: 10.1016/j.molcel.2007.04.020 - 46.
Soulard A, Cohen A, Hall MN. TOR signaling in invertebrates. Current Opinion in Cell Biology. 2009; 21 (6):825-836. DOI: 10.1016/j.ceb.2009.08.007 - 47.
Eltschinger S, Loewith R. TOR complexes and the maintenance of cellular homeostasis. Trends in Cell Biology. 2016; 26 (2):148-159. DOI: 10.1016/j.tcb.2015.10.003 - 48.
Peyroche G, Milkereit P, Bischler N, Tschochner H, Schultz P, Sentenac A, et al. The recruitment of RNA polymerase I on rDNA is mediated by the interaction of the A43 subunit with Rrn3. The EMBO Journal. 2000; 19 (20):5473-5482. DOI: 10.1093/emboj/19.20.5473 - 49.
Laferté A, Favry E, Sentenac A, Riva M, Carles C, Chédin S. The transcriptional activity of RNA polymerase I is a key determinant for the level of all ribosome components. Genes & Development. 2006; 20 (15):2030-2040. DOI: 10.1101/gad.386106 - 50.
Huber A, Bodenmiller B, Uotila A, Stahl M, Wanka S, Gerrits B, et al. Characterization of the rapamycin-sensitive phosphoproteome reveals that Sch9 is a central coordinator of protein synthesis. Genes & Development. 2009; 23 (16):1929-1943. DOI: 10.1101/gad.532109 - 51.
Jorgensen P, Rupeš I, Sharom JR, Schneper L, Broach JR, Tyers M. A dynamic transcriptional network communicates growth potential to ribosome synthesis and critical cell size. Genes & Development. 2004; 18 (20):2491-2505. DOI: 10.1101/gad.1228804 - 52.
Upadhya R, Lee JH, Willis IM. Maf1 is an essential mediator of diverse signals that repress RNA polymerase III transcription. Molecular Cell. 2002; 10 (6):1489-1494. DOI: 10.1016/s1097-2765(02)00787-6 - 53.
Oficjalska-Pham D, Harismendy O, Smagowicz WJ, Gonzalez de Peredo A, Boguta M, Sentenac A, et al. General repression of RNA polymerase III transcription is triggered by protein phosphatase type 2A-mediated dephosphorylation of Maf1. Molecular Cell. 2006; 22 (5):623-632. DOI: 10.1016/j.molcel.2006.04.008 - 54.
Roberts DN, Wilson B, Huff JT, Stewart AJ, Cairns BR. Dephosphorylation and genome-wide association of Maf1 with Pol III-transcribed genes during repression. Molecular Cell. 2006; 22 (5):633-644. DOI: 10.1016/j.molcel.2006.04.009 - 55.
Lee JH, Moir RD, Willis IM. Regulation of RNA polymerase III transcription involves SCH9-dependent and SCH9-independent branches of the target of rapamycin (TOR) pathway. The Journal of Biological Chemistry. 2009; 284 (19):12604-12608. DOI: 10.1074/jbc.C900020200 - 56.
Wei Y, Zheng XFS. Sch9 partially mediates TORC1 signaling to control ribosomal RNA synthesis. Cell Cycle. 2009; 8 (24):4085-4090. DOI: 10.4161/cc.8.24.10170 - 57.
Hughes Hallett JE, Luo X, Capaldi AP. State transitions in the TORC1 signaling pathway and information processing in Saccharomyces cerevisiae . Genetics. 2014;198 (2):773-786. DOI: 10.1534/genetics.114.168369 - 58.
Powers T, Walter P. Regulation of ribosome biogenesis by the rapamycin-sensitive TOR-signaling pathway in Saccharomyces cerevisiae . Molecular Biology of the Cell. 1999;10 (4):987-1000. DOI: 10.1091/mbc.10.4.987 - 59.
Boguta M. Maf1, a general negative regulator of RNA polymerase III in yeast. Biochimica et Biophysica Acta: Gene Regulatory Mechanisms. 2013; 1829 (3-4):376-384. DOI: 10.1016/j.bbagrm.2012.11.004 - 60.
Peisker K, Chiabudini M, Rospert S. The ribosome-bound Hsp70 homolog Ssb of Saccharomyces cerevisiae . Biochimica et Biophysica Acta: Molecular Cell Research. 2010;1803 (6):662-672. DOI: 10.1016/j.bbamcr.2010.03.005 - 61.
Bonawitz ND, Chatenay-Lapointe M, Pan Y, Shadel GS. Reduced TOR signaling extends chronological life span via increased respiration and upregulation of mitochondrial gene expression. Cell Metabolism. 2007; 5 (4):265-277. DOI: 10.1016/j.cmet.2007.02.009 - 62.
Caligaris M, Nicastro R, Hu Z, Tripodi F, Hummel JE, Pillet B, et al. Snf1/AMPK fine-tunes TORC1 signaling in response to glucose starvation. eLife. 2023; 12 :e84319. DOI: 10.7554/eLife.84319 - 63.
Carlson M, Osmond BC, Botstein D. Mutants of yeast defective in sucrose utilization. Genetics. 1981; 98 (1):25-40. DOI: 10.1093/genetics/98.1.25 - 64.
Celenza JL, Carlson M. A yeast gene that is essential for release from glucose repression encodes a protein kinase. Science. 1986; 233 (4769):1175-1180. DOI: 10.1126/science.3526554 - 65.
Hedbacker K, Carlson M. SNF1/AMPK pathways in yeast. Frontiers in Bioscience. 2008; 13 (13):2408. DOI: 10.2741/2854 - 66.
Hardie DG, Ross FA, Hawley SA. AMPK: A nutrient and energy sensor that maintains energy homeostasis. Nature Reviews Molecular Cell Biology. 2012; 13 (4):251-262. DOI: 10.1038/nrm3311 - 67.
Nehlin JO, Ronne H. Yeast MIG1 repressor is related to the mammalian early growth response and Wilms’ tumour finger proteins. The EMBO Journal. 1990; 9 (9):2891-2898. DOI: 10.1002/j.1460-2075.1990.tb07479.x - 68.
Treitel MA, Kuchin S, Carlson M. Snf1 protein kinase regulates phosphorylation of the Mig1 repressor in Saccharomyces cerevisiae . Molecular and Cellular Biology. 1998;18 (11):6273-6280. DOI: 10.1128/MCB.18.11.6273 - 69.
Schuller HJ, Entian KD. Extragenic suppressors of yeast glucose derepression mutants leading to constitutive synthesis of several glucose-repressible enzymes. Journal of Bacteriology. 1991; 173 (6):2045-2052. DOI: 10.1128/jb.173.6.2045-2052.1991 - 70.
Flick JS, Johnston M. Analysis of URS(G)-mediated glucose repression of the GAL1 promoter of Saccharomyces cerevisiae . Genetics. 1992;130 (2):295-304. DOI: 10.1093/genetics/130.2.295 - 71.
Vincent O, Townley R, Kuchin S, Carlson M. Subcellular localization of the Snf1 kinase is regulated by specific β subunits and a novel glucose signaling mechanism. Genes & Development. 2001; 15 (9):1104-1114. DOI: 10.1101/gad.879301 - 72.
Tu J, Carlson M. REG1 binds to protein phosphatase type 1 and regulates glucose repression in Saccharomyces cerevisiae . The EMBO Journal. 1995;14 (23):5939-5946. DOI: 10.1002/j.1460-2075.1995.tb00282.x - 73.
Sanz P, Alms GR, Haystead TAJ, Carlson M. Regulatory interactions between the Reg1-Glc7 protein phosphatase and the Snf1 protein kinase. Molecular and Cellular Biology. 2000; 20 (4):1321-1328. DOI: 10.1128/MCB.20.4.1321-1328.2000 - 74.
Rubenstein EM, McCartney RR, Zhang C, Shokat KM, Shirra MK, Arndt KM, et al. Access denied: Snf1 activation loop phosphorylation is controlled by availability of the phosphorylated threonine 210 to the PP1 phosphatase. The Journal of Biological Chemistry. 2008; 283 (1):222-230. DOI: 10.1074/jbc.M707957200 - 75.
Castermans D, Somers I, Kriel J, Louwet W, Wera S, Versele M, et al. Glucose-induced posttranslational activation of protein phosphatases PP2A and PP1 in yeast. Cell Research. 2012; 22 (6):1058-1077. DOI: 10.1038/cr.2012.20 - 76.
Ruiz A, Liu Y, Xu X, Carlson M. Heterotrimer-independent regulation of activation-loop phosphorylation of Snf1 protein kinase involves two protein phosphatases. Proceedings of the National Academy of Sciences of the United States of America. 2012; 109 (22):8652-8657. DOI: 10.1073/pnas.1206280109 - 77.
Ruiz A, Xu X, Carlson M. Ptc1 protein phosphatase 2C contributes to glucose regulation of SNF1/AMP-activated protein kinase (AMPK) in Saccharomyces cerevisiae . The Journal of Biological Chemistry. 2013;288 (43):31052-31058. DOI: 10.1074/jbc.M113.503763 - 78.
Özcan S, Dover J, Rosenwald AG, Wölfl S, Johnston M. Two glucose transporters in Saccharomyces cerevisiae are glucose sensors that generate a signal for induction of gene expression. Proceedings of the National Academy of Sciences of the United States of America. 1996;93 (22):12428-12432. DOI: 10.1073/pnas.93.22.12428 - 79.
Shashkova S, Welkenhuysen N, Hohmann S. Molecular communication: Crosstalk between the Snf1 and other signaling pathways. FEMS Yeast Research. 2015; 15 (4):fov026. DOI: 10.1093/femsyr/fov026 - 80.
Özcan S, Dover J, Johnston M. Glucose sensing and signaling by two glucose receptors in the yeast Saccharomyces cerevisiae . The EMBO Journal. 1998;17 (9):2566-2573. DOI: 10.1093/emboj/17.9.2566 - 81.
Özcan S, Johnston M. Three different regulatory mechanisms enable yeast hexose transporter (HXT) genes to be induced by different levels of glucose. Molecular and Cellular Biology. 1995; 15 (3):1564-1572. DOI: 10.1128/MCB.15.3.1564 - 82.
Tomás-Cobos L, Sanz P. Active Snf1 protein kinase inhibits expression of the Saccharomyces cerevisiae HXT1 glucose transporter gene. The Biochemical Journal. 2002;368 (2):657-663. DOI: 10.1042/BJ20020984 - 83.
Kim J-H, Polish J, Johnston M. Specificity and regulation of DNA binding by the yeast glucose transporter gene repressor Rgt1. Molecular and Cellular Biology. 2003; 23 (15):5208-5216. DOI: 10.1128/MCB.23.15.5208-5216.2003 - 84.
Lakshmanan J, Mosley AL, Özcan S. Repression of transcription by Rgt1 in the absence of glucose requires Std1 and Mth1. Current Genetics. 2003; 44 (1):19-25. DOI: 10.1007/s00294-003-0423-2 - 85.
Mosley AL, Lakshmanan J, Aryal BK, Özcan S. Glucose-mediated phosphorylation converts the transcription factor Rgt1 from a repressor to an activator. The Journal of Biological Chemistry. 2003; 278 (12):10322-10327. DOI: 10.1074/jbc.M212802200 - 86.
Polish JA, Kim JH, Johnston M. How the Rgt1 transcription factor of Saccharomyces cerevisiae is regulated by glucose. Genetics. 2005;169 (2):583-594. DOI: 10.1534/genetics.104.034512 - 87.
Moriya H, Johnston M. Glucose sensing and signaling in Saccharomyces cerevisiae through the Rgt2 glucose sensor and casein kinase I. Proceedings of the National Academy of Sciences of the United States of America. 2004;101 (6):1572-1577. DOI: 10.1073/pnas.0305901101 - 88.
Flick KM, Spielewoy N, Kalashnikova TI, Guaderrama M, Zhu Q , Chang H-C, et al. Grr1-dependent inactivation of Mth1 mediates glucose-induced dissociation of Rgt1 from HXT gene promoters. Molecular Biology of the Cell. 2003; 14 (8):3230-3241. DOI: 10.1091/mbc.e03-03-0135 - 89.
Kim JH, Brachet V, Moriya H, Johnston M. Integration of transcriptional and posttranslational regulation in a glucose signal transduction pathway in Saccharomyces cerevisiae . Eukaryotic Cell. 2006;5 (1):167-173. DOI: 10.1128/EC.5.1.167-173.2006 - 90.
Zaman S, Lippman SI, Schneper L, Slonim N, Broach JR. Glucose regulates transcription in yeast through a network of signaling pathways. Molecular Systems Biology. 2009; 5 (245):1-14. DOI: 10.1038/msb.2009.2 - 91.
Gagiano M, Bauer FF, Pretorius IS. The sensing of nutritional status and the relationship to filamentous growth in Saccharomyces cerevisiae . FEMS Yeast Research. 2002;2 (4):433-470. DOI: 10.1111/j.1567-1364.2002.tb00114.x - 92.
Lackner LL. Shaping the dynamic mitochondrial network. BMC Biology. 2014; 12 (1):35. DOI: 10.1186/1741-7007-12-35 - 93.
Tsuboi T, Viana MP, Xu F, Yu J, Chanchani R, Arceo XG, et al. Mitochondrial volume fraction and translation duration impact mitochondrial mRNA localization and protein synthesis. eLife. 2020; 9 (529289):e57814. DOI: 10.7554/eLife.57814 - 94.
Chacinska A, Koehler CM, Milenkovic D, Lithgow T, Pfanner N. Importing mitochondrial proteins: Machineries and mechanisms. Cell. 2009; 138 (4):628-644. DOI: 10.1016/j.cell.2009.08.005 - 95.
Wiedemann N, Pfanner N. Mitochondrial machineries for protein import and assembly. Annual Review of Biochemistry. 2017; 86 (1):685-714. DOI: 10.1146/annurev-biochem-060815-014352 - 96.
Avendaño-Monsalve MC, Ponce-Rojas JC, Funes S. From cytosol to mitochondria: The beginning of a protein journey. Biological Chemistry. 2020; 401 (6-7):645-661. DOI: 10.1515/hsz-2020-0110 - 97.
Garcia M, Darzacq X, Delaveau T, Jourdren L, Singer RH, Jacq C. Mitochondria-associated yeast mRNAs and the biogenesis of molecular complexes. Fox T, editor. Molecular Biology of the Cell. 2007; 18 (2):362-368. DOI: 10.1091/mbc.e06-09-0827 - 98.
Marc P, Margeot A, Devaux F, Blugeon C, Corral-Debrinski M, Jacq C. Genome-wide analysis of mRNAs targeted to yeast mitochondria. EMBO Reports. 2002; 3 (2):159-164. DOI: 10.1093/embo-reports/kvf025 - 99.
Lesnik C, Golani-Armon A, Arava Y. Localized translation near the mitochondrial outer membrane: An update. RNA Biology. 2015; 12 (8):801-809. DOI: 10.1080/15476286.2015.1058686 - 100.
Sylvestre J, Vialette S, Corral Debrinski M, Jacq C. Long mRNAs coding for yeast mitochondrial proteins of prokaryotic origin preferentially localize to the vicinity of mitochondria. Genome Biology. 2003; 4 :R44. DOI: 10.1186/gb-2003-4-7-r44 - 101.
Williams CC, Jan CH, Weissman JS. Targeting and plasticity of mitochondrial proteins revealed by proximity-specific ribosome profiling. Science. 2014; 346 (6210):748-751. DOI: 10.1126/science.1257522 - 102.
Gold VA, Chroscicki P, Bragoszewski P, Chacinska A. Visualization of cytosolic ribosomes on the surface of mitochondria by electron cryotomography. EMBO Reports. 2017; 18 (10):1786-1800. DOI: 10.15252/embr.201744261 - 103.
Cavalier-Smith T. Origin of mitochondria by intracellular enslavement of a photosynthetic purple bacterium. Proceedings of the Royal Society B: Biological Sciences. 2006; 273 (1596):1943-1952. DOI: 10.1098/rspb.2006.3531 - 104.
Bonitz SG, Berlani R, Coruzzi G, Li M, Macino G, Nobrega FG, et al. Codon recognition rules in yeast mitochondria. Proceedings of the National Academy of Sciences of the United States of America. 1980; 77 (6I):3167-3170. DOI: 10.1073/pnas.77.6.3167 - 105.
Salinas-Giegé T, Giegé R, Giegé P. tRNA biology in mitochondria. International Journal of Molecular Sciences. 2015; 16 (3):4518-4559. DOI: 10.3390/ijms16034518 - 106.
Scheffler IE, De La Cruz BJ, Prieto S. Control of mRNA turnover as a mechanism of glucose repression in Saccharomyces cerevisiae . The International Journal of Biochemistry & Cell Biology. 1998;30 (11):1175-1193. DOI: 10.1016/s1357-2725(98)00086-7 - 107.
Miller MA, Russo J, Fischer AD, Leban FAL, Olivas WM. Carbon source-dependent alteration of Puf3p activity mediates rapid changes in the stabilities of mRNAs involved in mitochondrial function. Nucleic Acids Research. 2014; 42 (6):3954-3970. DOI: 10.1093/nar/gkt1346 - 108.
Foat BC, Houshmandi SS, Olivas WM, Bussemaker HJ. Profiling condition-specific, genome-wide regulationof mRNA stability in yeast. Proceedings of the National Academy of Sciences of the United States of America. 2005; 102 (49):17675-17680. DOI: 10.1073/pnas.0503803102 - 109.
Endo T, Yamano K, Kawano S. Structural insight into the mitochondrial protein import system. Biochimica et Biophysica Acta (BBA) - Biomembranes. 2011; 1808 (3):955-970. DOI: 10.1016/j.bbamem.2010.07.018 DOI: 10.1016/j.bbamem.2010.07.018 - 110.
Priesnitz C, Becker T. Pathways to balance mitochondrial translation and protein import. Genes & Development. 2018; 32 (19-20):1285-1296. DOI: 10.1101/gad.316547.118 - 111.
Couvillion MT, Soto IC, Shipkovenska G, Churchman LS. Synchronized mitochondrial and cytosolic translation programs. Nature. 2016; 533 (7604):499-503. DOI: 10.1038/nature18015 - 112.
Wang X, Chen XJ. A cytosolic network suppressing mitochondria-mediated proteostatic stress and cell death. Nature. 2015; 524 (7566):481-484. DOI: 10.1038/nature14859 - 113.
Andréasson C, Ott M, Büttner S. Mitochondria orchestrate proteostatic and metabolic stress responses. EMBO Reports. 2019; 20 (10):e47865. DOI: 10.15252/embr.201947865 - 114.
Lill R, Mühlenhoff U. Maturation of iron-sulfur proteins in eukaryotes: Mechanisms, connected processes, and diseases. Annual Review of Biochemistry. 2008; 77 (1):669-700. DOI: 10.1146/annurev.biochem.76.052705.162653 - 115.
Attardi G, Schatz G. Biogenesis of mitochondria. Annual Review of Cell Biology. 1988; 4 :289-333. DOI: 10.1146/annurev.cb.04.110188.001445 - 116.
Malina C, Larsson C, Nielsen J. Yeast mitochondria: An overview of mitochondrial biology and the potential of mitochondrial systems biology. FEMS Yeast Research. 2018; 18 (5):foy040. DOI: 10.1093/femsyr/foy040 - 117.
Gancedo JM. Yeast carbon catabolite repression. Microbiology and Molecular Biology Reviews. 1998; 62 (2):334-361. DOI: 10.1128/MMBR.62.2.334-361.1998 - 118.
Sharma AK, Pallesen LJ, Spang RJ, Walden WE. Cytosolic iron-sulfur cluster assembly (CIA) system: Factors, mechanism, and relevance to cellular iron regulation. The Journal of Biological Chemistry. 2010; 285 (35):26745-26751. DOI: 10.1074/jbc.R110.122218 - 119.
Ramos-Alonso L, Romero AM, Martínez-Pastor MT, Puig S. Iron regulatory mechanisms in Saccharomyces cerevisiae . Frontiers in Microbiology. 2020;11 :582830. DOI: 10.3389/fmicb.2020.582830 - 120.
Romero AM, Martínez-Pastor MT, Puig S. Iron in translation: From the beginning to the end. Microorganisms. 2021; 9 (5):1058. DOI: 10.3390/microorganisms9051058 - 121.
Puig S, Vergara SV, Thiele DJ. Cooperation of two mRNA-binding proteins drives metabolic adaptation to iron deficiency. Cell Metabolism. 2008; 7 (6):555-564. DOI: 10.1016/j.cmet.2008.04.010 - 122.
Yamaguchi-Iwai Y, Dancis A, Klausner RD. AFT1: A mediator of iron regulated transcriptional control in Saccharomyces cerevisiae . The EMBO Journal. 1995;14 (6):1231-1239. DOI: 10.1002/j.1460-2075.1995.tb07106.x - 123.
Haurie V, Boucherie H, Sagliocco F. The Snf1 protein kinase controls the induction of genes of the iron uptake pathway at the diauxic shift in Saccharomyces cerevisiae . The Journal of Biological Chemistry. 2003;278 (46):45391-45396. DOI: 10.1074/jbc.M307447200 - 124.
Philpott CC, Protchenko O. Response to iron deprivation in Saccharomyces cerevisiae . Eukaryotic Cell. 2008;7 (1):20-27. DOI: 10.1128/EC.00354-07 - 125.
Lill R, Mühlenhoff U. Iron-sulfur protein biogenesis in eukaryotes: Components and mechanisms. Annual Review of Cell and Developmental Biology. 2006; 22 :457-486. DOI: 10.1146/annurev.cellbio.22.010305.104538 - 126.
Holmes-Hampton GP, Jhurry ND, McCormick SP, Lindahl PA, Jo WJ, Hyoun JH, et al. Response to iron deprivation in Saccharomyces cerevisiae . Biochemistry. 2008;10 (1):105-114. DOI: 10.1128/EC.00354-07 - 127.
Martins TS, Costa V, Pereira C. Signaling pathways governing iron homeostasis in budding yeast. Molecular Microbiology. 2018; 109 (4):422-432. DOI: 10.1111/mmi.14009 - 128.
den Ridder M, Daran-Lapujade P, Pabst M. Shot-gun proteomics: Why thousands of unidentified signals matter. FEMS Yeast Research. 2020; 20 (1):foz088. DOI: 10.1093/femsyr/foz088 - 129.
Daran-Lapujade P, Rossell S, Van Gulik WM, Luttik MAH, De Groot MJL, Slijper M, et al. The fluxes through glycolytic enzymes in Saccharomyces cerevisiae are predominantly regulated at posttranscriptional levels. Proceedings of the National Academy of Sciences of the United States of America. 2007;104 (40):15753-15758. DOI: 10.1073/pnas.0707476104 - 130.
Soares Rodrigues CI, den Ridder M, Pabst M, Gombert AK, Wahl SA. Comparative proteome analysis of different Saccharomyces cerevisiae strains during growth on sucrose and glucose. Scientific Reports. 2023;13 (1):2126. DOI: 10.1038/s41598-023-29172-0 - 131.
Kolkman A, Olsthoorn MMA, Heeremans CEM, Heck AJR, Slijper M. Comparative proteome analysis of Saccharomyces cerevisiae grown in chemostat cultures limited for glucose or ethanol. Molecular & Cellular Proteomics. 2005;4 (1):1-11. DOI: 10.1074/mcp.M400087-MCP200 - 132.
Garcia-Albornoz M, Holman SW, Antonisse T, Daran-Lapujade P, Teusink B, Beynon RJ, et al. A proteome-integrated, carbon source dependent genetic regulatory network in: Saccharomyces cerevisiae . Molecular Oral Microbiology. 2020;16 (1):59-72. DOI: 10.1039/C9MO00136K - 133.
Paulo JA, O’Connell JD, Everley RA, O’Brien J, Gygi MA, Gygi SP. Quantitative mass spectrometry-based multiplexing compares the abundance of 5000 S. cerevisiae proteins across 10 carbon sources. Journal of Proteomics. 2016; 148 :85-93. DOI: 10.1016/j.jprot.2016.07.005 - 134.
Paulo JA, O’Connell JD, Gaun A, Gygi SP. Proteome-wide quantitative multiplexed profiling of protein expression: Carbon-source dependency in Saccharomyces cerevisiae . Molecular Biology of the Cell. 2015;26 (22):4063-4074. DOI: 10.1091/mbc.E15-07-0499 - 135.
Pichon X, Wilson LA, Stoneley M, Bastide A, King HA, Somers J, et al. RNA binding protein/RNA element interactions and the control of translation. Current Protein & Peptide Science. 2012; 13 (4):294-304. DOI: 10.2174/138920312801619475 - 136.
Simpson CE, Ashe MP. Adaptation to stress in yeast: To translate or not? Biochemical Society Transactions. 2012; 40 (4):794-799. DOI: 10.1042/BST20120078 - 137.
Ashe MP, De LSK, Sachs AB. Glucose depletion rapidly inhibits translation initiation in yeast. Molecular Biology of the Cell. 2000; 11 :833-848. DOI: 10.1091/mbc.11.3.833 - 138.
Buchan JR, Nissan T, Parker R. Analyzing P-bodies and stress granules in Saccharomyces cerevisiae . Methods in Enzymology. 2010;470 :619-640. DOI: 10.1016/S0076-6879(10)70025-2 - 139.
Kedersha N, Stoecklin G, Ayodele M, Yacono P, Lykke-Andersen J, Fitzler MJ, et al. Stress granules and processing bodies are dynamically linked sites of mRNP remodeling. The Journal of Cell Biology. 2005; 169 (6):871-884. DOI: 10.1083/jcb.200502088 - 140.
Xing W, Muhlrad D, Parker R, Rosen MK. A quantitative inventory of yeast P body proteins reveals principles of composition and specificity. eLife. 2020; 9 :e56525. DOI: 10.7554/eLife.56525 - 141.
Balagopal V, Fluch L, Nissan T. Ways and means of eukaryotic mRNA decay. Biochimica et Biophysica Acta: Gene Regulatory Mechanisms. 2012; 1819 (6):593-603. DOI: 10.1016/j.bbagrm.2012.01.001 - 142.
Balagopal V, Parker R. Polysomes, P bodies and stress granules: States and fates of eukaryotic mRNAs. Current Opinion in Cell Biology. 2009; 21 (3):403-408. DOI: 10.1016/j.ceb.2009.03.005 - 143.
Luo Y, Na Z, Slavoff SA. P-bodies: Composition, properties, and functions. Biochemistry. 2018; 57 (17):2424-2431. DOI: 10.1021/acs.biochem.7b01162 - 144.
Decker CJ, Parker R. P-bodies and stress granules: Possible roles in the control of translation and mRNA degradation. Cold Spring Harbor Perspectives in Biology. 2012; 4 (9):a012286. DOI: 10.1101/cshperspect.a012286 - 145.
Drummond SP, Hildyard J, Firczuk H, Reamtong O, Li N, Kannambath S, et al. Diauxic shift-dependent relocalization of decapping activators Dhh1 and Pat1 to polysomal complexes. Nucleic Acids Research. 2011; 39 (17):7764-7774. DOI: 10.1093/nar/gkr474 - 146.
Phizicky EM, Hopper AK. The life and times of a tRNA. RNA. 2023; 29 (7):898-957. DOI: 10.1261/rna.079620.123 - 147.
Warner JR. The economics of ribosome biosynthesis in yeast. Trends in Biochemical Sciences. 1999; 24 (11):437-440. DOI: 10.1016/s0968-0004(99)01460-7 - 148.
Quax TEF, Claassens NJ, Söll D, van der Oost J. Codon bias as a means to fine-tune gene expression. Molecular Cell. 2015; 59 (2):149-161. DOI: 10.1016/j.molcel.2015.05.035 - 149.
Chan PP, Lowe TM. GtRNAdb 2.0: An expanded database of transfer RNA genes identified in complete and draft genomes. Nucleic Acids Research. 2016; 44 (D1):D184-D189. DOI: 10.1093/nar/gkv1309 - 150.
Hani J, Feldmann H. tRNA genes and retroelements in the yeast genome. Nucleic Acids Research. 1998; 26 (3):689-696. DOI: 10.1093/nar/26.3.689 - 151.
Guimarães AR, Correia I, Sousa I, Oliveira C, Moura G, Bezerra AR, et al. tRNAs as a driving force of genome evolution in yeast. Frontiers in Microbiology. 2021; 12 :634004. DOI: 10.3389/fmicb.2021.634004 - 152.
Tavares JF, Davis NK, Poim A, Reis A, Kellner S, Sousa I, et al. tRNA-modifying enzyme mutations induce codon-specific mistranslation and protein aggregation in yeast. RNA Biology. 2021; 18 (4):563-575. DOI: 10.1080/15476286.2020.1819671 - 153.
Edwards AM, Addo MA, Dos Santos PC. Extracurricular functions of tRNA modifications in microorganisms. Genes (Basel). 2020; 11 (8):907. DOI: 10.3390/genes11080907 - 154.
Tarassov I, Entelis N, Martin RP. Mitochondrial import of a cytoplasmic lysine-tRNA in yeast is mediated by cooperation of cytoplasmic and mitochondrial lysyl-tRNA synthetases. The EMBO Journal. 1995; 14 (14):3461-3471. DOI: 10.1002/j.1460-2075.1995.tb07352.x - 155.
Rinehart J, Krett B, Rubio MAT, Alfonzo JD, Söll D. Saccharomyces cerevisiae imports the cytosolic pathway for Gln-tRNA synthesis into the mitochondrion. Genes & Development. 2005;19 (5):583-592. DOI: 10.1101/gad.1269305 - 156.
Rubio MAT, Hopper AK. Transfer RNA travels from the cytoplasm to organelles. Wiley Interdisciplinary Reviews RNA. 2011; 2 (6):802-817. DOI: 10.1002/wrna.93 - 157.
Bruch A, Laguna T, Butter F, Schaffrath R, Klassen R. Misactivation of multiple starvation responses in yeast by loss of tRNA modifications. Nucleic Acids Research. 2020; 48 (13):7307-7320. DOI: 10.1093/nar/gkaa455 - 158.
Hayashi S, Mori S, Suzuki T, Suzuki T, Yoshihisa T. Impact of intron removal from tRNA genes on Saccharomyces cerevisiae . Nucleic Acids Research. 2019;47 (11):5936-5949. DOI: 10.1093/nar/gkz270 - 159.
Baldi MI, Mattoccia E, Bufardeci E, Fabbri S, Tocchini-Valentini GP. Participation of the intron in the reaction catalyzed by the xenopus tRNA splicing endonuclease. Science. 1992; 255 (5050):1404-1408. DOI: 10.1126/science.1542788 - 160.
Negri EDN, Fabbri S, Bufardeci E, Baldi MI, Attardi DG, Mattoccia E, et al. The eucaryal tRNA splicing endonuclease recognizes a tripartite set of RNA elements. Cell. 1997; 89 (6):859-866. DOI: 10.1016/s0092-8674(00)80271-8 - 161.
Yoshihisa T, Yunoki-Esaki K, Ohshima C, Tanaka N, Endo T. Possibility of cytoplasmic pre-tRNA splicing: The yeast tRNA splicing endonuclease mainly localizes on the mitochondria. Molecular Biology of the Cell. 2003; 14 (8):3266-3279. DOI: 10.1091/mbc.E02-11-0757 - 162.
Hopper AK, Nostramo RT. tRNA processing and subcellular trafficking proteins multitask in pathways for other RNAs. Frontiers in Genetics. 2019; 10 (FEB):1-14. DOI: 10.3389/fgene.2019.00096 - 163.
Huang HY, Hopper AK. Multiple layers of stress-induced regulation in tRNA biology. Life. 2016; 6 (2):13-16. DOI: 10.3390/life6020016 - 164.
Holcik M, Sonenberg N. Translational control in stress and apoptosis. Nature Reviews Molecular Cell Biology. 2005; 6 (4):318-327. DOI: 10.1038/nrm1618 - 165.
Chatterjee K, Nostramo RT, Wan Y, Hopper AK. tRNA dynamics between the nucleus, cytoplasm and mitochondrial surface: Location, location, location. Biochimica et Biophysica Acta: Gene Regulatory Mechanisms. 2018; 1861 (4):373-386. DOI: 10.1016/j.bbagrm.2017.11.007 - 166.
Chu H-Y, Hopper AK. Genome-wide investigation of the role of the tRNA nuclear-cytoplasmic trafficking pathway in regulation of the yeast Saccharomyces cerevisiae transcriptome and proteome. Molecular and Cellular Biology. 2013;33 (21):4241-4254. DOI: 10.1128/MCB.00785-13 - 167.
Whitney ML, Hurto RL, Shaheen HH, Hopper AK. Rapid and reversible nuclear accumulation of cytoplasmic tRNA in response to nutrient availability. Fox T, editor. Molecular Biology of the Cell. 2007; 18 (7):2678-2686. DOI: 10.1091/mbc.e07-01-0006 - 168.
Karkusiewicz I, Turowski TW, Graczyk D, Towpik J, Dhungel N, Hopper AK, et al. Maf1 protein, repressor of RNA polymerase III, indirectly affects tRNA processing. The Journal of Biological Chemistry. 2011; 286 (45):39478-39488. DOI: 10.1074/jbc.M111.253310 - 169.
Phizicky EM, Hopper AK. tRNA biology charges to the front. Genes & Development. 2010; 24 (17):1832-1860. DOI: 10.1101/gad.1956510 - 170.
Yoshihisa T. Handling tRNA introns, archaeal way and eukaryotic way. Frontiers in Genetics. 2014; 5 (JUL):1-16. DOI: 10.3389/fgene.2014.00213 - 171.
Fujishima K, Kanai A. tRNA gene diversity in the three domains of life. Frontiers in Genetics. 2014; 5 :142. DOI: 10.3389/fgene.2014.00142 - 172.
Schmidt CA, Matera AG. tRNA introns: Presence, processing, and purpose. WIREs RNA [Internet]. 2020; 11 (3):e1583. DOI: 10.1002/wrna.1583 - 173.
Walters R, Parker R. Is there quality control of localized mRNAs? The Journal of Cell Biology. 2014; 204 (6):863-868. DOI: 10.1083/jcb.201401059 - 174.
Quenault T, Lithgow T, Traven A. PUF proteins: Repression, activation and mRNA localization. Trends in Cell Biology. 2011; 21 (2):104-112. DOI: 10.1016/j.tcb.2010.09.013 - 175.
Crawford RA, Pavitt GD. Translational regulation in response to stress in Saccharomyces cerevisiae . Yeast. 2019;36 (1):5-21. DOI: 10.1002/yea.3349 - 176.
Gerber AP, Herschlag D, Brown PO. Extensive association of functionally and cytotopically related mRNAs with Puf family RNA-binding proteins in yeast. PLoS Biology. 2004; 2 (3):E79. DOI: 10.1371/journal.pbio.0020079 - 177.
Hogan DJ, Riordan DP, Gerber AP, Herschlag D, Brown PO. Diverse RNA-binding proteins interact with functionally related sets of RNAs, suggesting an extensive regulatory system. PLoS Biology. 2008; 6 (10):2297-2313. DOI: 10.1371/journal.pbio.0060255 - 178.
Freeberg MA, Han T, Moresco JJ, Kong A, Yang YC, Lu ZJ, et al. Pervasive and dynamic protein binding sites of the mRNA transcriptome in Saccharomyces cerevisiae . Genome Biology. 2013;14 (2):R13. DOI: 10.1186/gb-2013-14-2-r13 - 179.
Kershaw CJ, Costello JL, Talavera D, Rowe W, Castelli LM, Sims PFG, et al. Integrated multi-omics analyses reveal the pleiotropic nature of the control of gene expression by Puf3p. Scientific Reports. 2015; 5 :15518. DOI: 10.1038/srep15518 - 180.
Edwards TA, Pyle SE, Wharton RP, Aggarwal AK. Structure of Pumilio reveals similarity between RNA and peptide binding motifs. Cell. 2001; 105 (2):281-289. DOI: 10.1016/s0092-8674(01)00318-x - 181.
Wang X, Zamore PD, Tanaka Hall TM. Crystal structure of a Pumilio homology domain. Molecular Cell. 2001; 7 (4):855-865. DOI: 10.1016/s1097-2765(01)00229-5 - 182.
Zhu D, Stumpf CR, Krahn JM, Wickens M, Tanaka Hall TM. A 5′ cytosine binding pocket in Puf3p specifies regulation of mitochondrial mRNAs. Proceedings of the National Academy of Sciences of the United States of America. 2009; 106 (48):20192-20197. DOI: 10.1073/pnas.0812079106 - 183.
Cheong CG, Tanaka Hall TM. Engineering RNA sequence specificity of Pumilio repeats. Proceedings of the National Academy of Sciences of the United States of America. 2006; 103 (37):13635-13639. DOI: 10.1073/pnas.0606294103 - 184.
Koh YY, Opperman L, Stumpf C, Mandan A, Keles S, Wickens M. A single C. elegans PUF protein binds RNA in multiple modes. RNA. 2009; 15 (6):1090-1099. DOI: 10.1261/rna.1545309 - 185.
Miller MT, Higgin JJ, Tanaka Hall TM. Basis of altered RNA-binding specificity by PUF proteins revealed by crystal structures of yeast Puf4p. Nature Structural & Molecular Biology. 2008; 15 (4):397-402. DOI: 10.1038/nsmb.1390 - 186.
Opperman L, Hook B, DeFino M, Bernstein DS, Wickens M. A single spacer nucleotide determines the specificities of two mRNA regulatory proteins. Nature Structural & Molecular Biology. 2005; 12 (11):945-951. DOI: 10.1038/nsmb1010 - 187.
Varshney U, Lee CP, RajBhandary UL. Direct analysis of aminoacylation levels of tRNAs in vivo: Application to studying recognition of Escherichia coli initiator tRNA mutants by glutaminyl-tRNA synthetase. The Journal of Biological Chemistry. 1991;266 (36):24712-24718 - 188.
Wang Y, Opperman L, Wickens M, Tanaka Hall TM. Structural basis for specific recognition of multiple mRNA targets by a PUF regulatory protein. Proceedings of the National Academy of Sciences of the United States of America. 2009; 106 (48):20186-20191. DOI: 10.1073/pnas.0812076106 - 189.
Houshmandi SS, Olivas WM. Yeast Puf3 mutants reveal the complexity of Puf-RNA binding and identify a loop required for regulation of mRNA decay. RNA. 2005; 11 (11):1655-1666. DOI: 10.1261/rna.2168505 - 190.
Jackson JS, Houshmandi SS, Leban FL, Olivas WM. Recruitment of the Puf3 protein to its mRNA target for regulation of mRNA decay in yeast. RNA. 2004; 10 (10):1625-1636. DOI: 10.1261/rna.7270204 - 191.
Olivas W, Parker R. The Puf3 protein is a transcript-specific regulator of mRNA degradation in yeast. The EMBO Journal. 2000; 19 (23):6602-6611. DOI: 10.1093/emboj/19.23.6602 - 192.
Riordan DP, Herschlag D, Brown PO. Identification of RNA recognition elements in the Saccharomyces cerevisiae transcriptome. Nucleic Acids Research. 2011;39 (4):1501-1509. DOI: 10.1093/nar/gkq920 - 193.
Bernstein D, Hook B, Hajarnavis A, Opperman L, Wickens M. Binding specificity and mRNA targets of a C. elegans PUF protein, FBF-1. RNA. 2005; 11 (4):447-458. DOI: 10.1261/rna.7255805 - 194.
Wickens M, Bernstein DS, Kimble J, Parker R. A PUF family portrait: 3’UTR regulation as a way of life. Trends in Genetics. 2002; 18 (3):150-157. DOI: 10.1016/S0168-9525(01)02616-6 - 195.
Campbell ZT, Valley CT, Wickens M. A protein-RNA specificity code enables targeted activation of an endogenous human transcript. Nature Structural & Molecular Biology. 2014; 21 (8):732-738. DOI: 10.1038/nsmb.2847 - 196.
Zhou W, Melamed D, Banyai G, Meyer C, Tuschl T, Wickens M, et al. Expanding the binding specificity for RNA recognition by a PUF domain. Nature Communications. 2021; 12 :5107. DOI: 10.1038/s41467-021-25433-6 - 197.
Gupta YK, Nair DT, Wharton RP, Aggarwal AK. Structures of human Pumilio with noncognate RNAs reveal molecular mechanisms for binding promiscuity. Structure. 2008; 16 (4):549-557. DOI: 10.1016/j.str.2008.01.006 - 198.
Lee CD, Tu BP. Glucose-regulated phosphorylation of the PUF protein Puf3 regulates the translational fate of its bound mRNAs and association with RNA granules. Cell Reports. 2015; 11 (10):1638-1650. DOI: 10.1016/j.celrep.2015.05.014 - 199.
García-Rodríguez LJ, Gay AC, Pon LA. Puf3p, a Pumilio family RNA binding protein, localizes to mitochondria and regulates mitochondrial biogenesis and motility in budding yeast. The Journal of Cell Biology. 2007; 176 (2):197-207. DOI: 10.1083/jcb.200606054 - 200.
Rowe W, Kershaw CJ, Castelli LM, Costello JL, Ashe MP, Grant CM, et al. Puf3p induces translational repression of genes linked to oxidative stress. Nucleic Acids Research. 2014; 42 (2):1026-1041. DOI: 10.1093/nar/gkt948 - 201.
Chatenay-Lapointe M, Shadel GS. Repression of mitochondrial translation, respiration and a metabolic cycle-regulated gene, SLF1, by the yeast Pumilio-family protein Puf3p. PLoS One. 2011; 6 (5):e20441. DOI: 10.1371/journal.pone.0020441 - 202.
Gupta I, Clauder-Münster S, Klaus B, Järvelin AI, Aiyar RS, Benes V, et al. Alternative polyadenylation diversifies post-transcriptional regulation by selective RNA-protein interactions. Molecular Systems Biology. 2014; 10 (2):1-11. DOI: 10.1002/msb.135068 - 203.
Lee D, Ohn T, Chiang Y-C, Quigley G, Yao G, Liu Y, et al. PUF3 acceleration of deadenylation in vivo can operate independently of CCR4 activity, possibly involving effects on the PAB1–mRNP structure. Journal of Molecular Biology. 2010; 399 (4):562-575. DOI: 10.1016/j.jmb.2010.04.034 - 204.
Wang Z, Sun X, Wee J, Guo X, Gu Z. Novel insights into global translational regulation through Pumilio family RNA-binding protein Puf3p revealed by ribosomal profiling. Current Genetics. 2019; 65 (1):201-212. DOI: 10.1007/s00294-018-0862-4 - 205.
Bhondeley M, Liu Z. Mitochondrial biogenesis is positively regulated by casein kinase I Hrr25 through phosphorylation of Puf3 in Saccharomyces cerevisiae . Genetics. 2020;215 (2):463-482. DOI: 10.1534/genetics.120.303191 - 206.
Forsburg SL, Guarente L. Identification and characterization of HAP4: A third component of the CCAAT-bound HAP2/HAP3 heteromer. Genes & Development. 1989; 3 (8):1166-1178. DOI: 10.1101/gad.3.8.1166